MonaLisa Documentation page

This is the documentation page of the MonaLisa program, a multilevel optimizer developed in the Quantum Chemistry group of Professor Joachim Sauer at Humbold-Universität zu Berlin.
MonaLisa is an optimizer for molecules and extended systems.
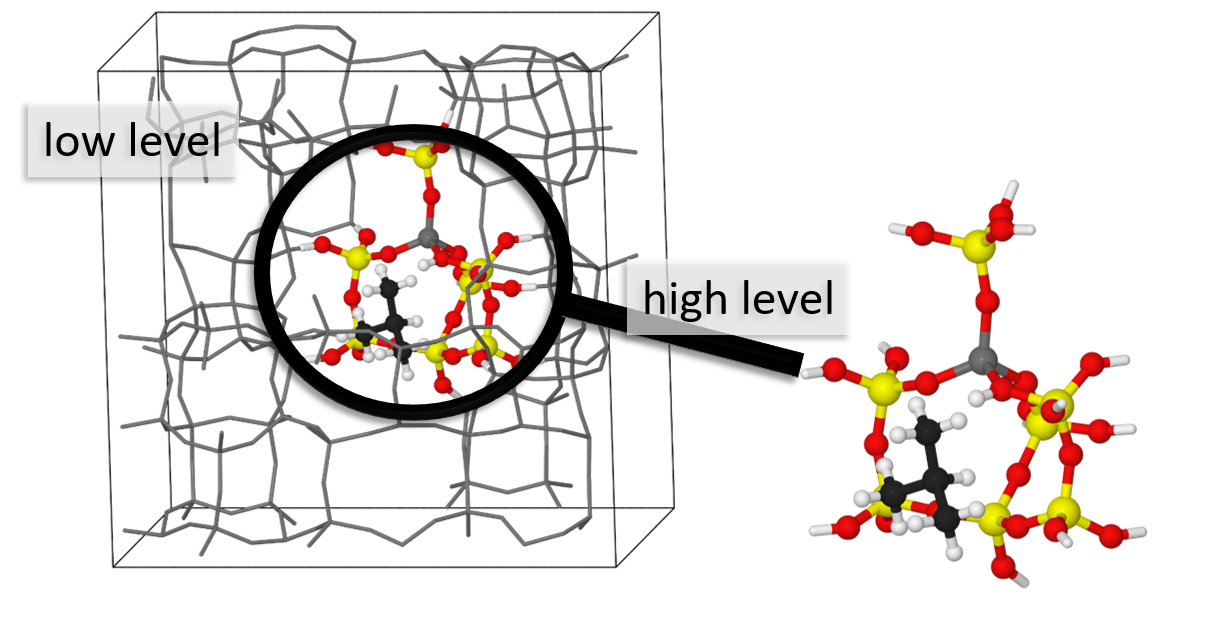
Its key feature is a generalized subtractive QM:QM:... scheme, also called multi-level embedding. In this approach a system (molecule) can be divided into arbitrary number of sub-parts which can be treated using different quantum chemistry methods, implemented in external programs. MonaLisa drives the optimization of the supersystem, but it can also do many more.
MonaLisa consists of the Runners and Interfaces
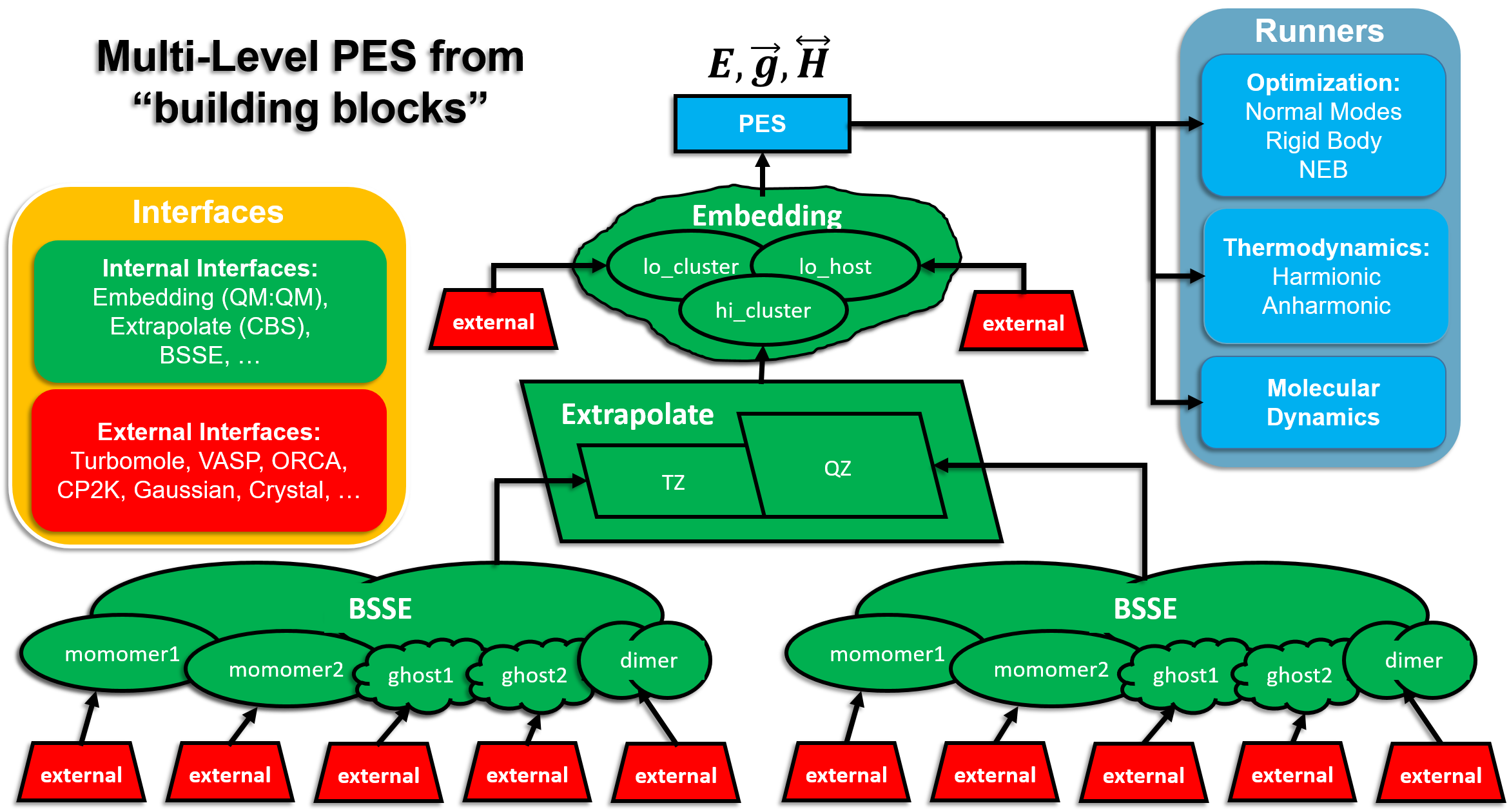
Runners
Runners are responsible for exploring the potential energy surface (PES). In other words they manipulate the structure of the molecule and analyze the molecular properties provided by Interfaces
Runner Types
Interfaces
MonaLisa use the eternal molecular modeling programs to obtain the information about the molecular properties of the investigated molecular system. The interaction between MonaLisa and external programs proceed via so called External Interfaces which currently can handle the following programs. The data provided by several interfaces may be analyzed by Internal Interfaces, which provide the hybrid molecular properties (energies, gradients, etc.)