- Course preparation: R basics
- Why R?
- Operators
- Objects
- Data Types and Data Structures
- Vectors
- Matrices and Arrays
- Data Frames and Lists
- Indexing
- Missing Values
- Flow Control
- Loops
- if-statements
- Functions
- Data Types
- Tutorials
- Coding style
- Help!
- Setting up a project in RStudio
- Test for Session 3:
terra-package
Course preparation: R basics
Here, we aim to provide a brief overview on the basics of R, which is supposed to refresh your knowledge and help you to identify topics worth revisiting. You are encouraged to refresh your knowledge from the statistic module as well as learn from online resources we list at the bottom of this page.
Why R?
R is a programming language and open source software environment for statistical computing and graphics. The R language is widely used among statisticians and data miners for developing statistical software and data analysis. It was developed by Ross Ihaka and Robert Gentleman at the University of Auckland, New Zealand. The name R originates the first names of the two authors and refers to the programming language S. The project was conceived in 1992, with an initial version released in 1995 and a stable beta version in 2000.
Learning R has tons of advantages. It is a great starting point for
those eager to learn programming. R offers increasingly specialized
tools for data wrangling, statistical analyses, and visualization. The
CRAN package repository currently features >13,000 packages serving a
variety of purposes, e.g. data manipulation (tidyr,
dplyr, caret), visualization
(ggplot2, ggmap, rasterVis), and
geodata handling (raster, rgdal,
sp, sf). You will notice that a huge share of
figures in scientific publications was produced using R. The R community
is huge, and offers great support. R is extremely popular in science
& industry, so a proficiency in R opens a wide array of job
opportunities. Everything is free and open source.
Operators
R is an interpreted programming language which means users can execute commands directly in the console without having to compile sourcecode. You can use R as a simple calculator using arithmetic operators and mathematical functions which are then evaluated directly. For example:
# A simple calculation
2*(1+4)
# abs() returns absolute values
abs(-2)
# sqrt() returns the squareroot
sqrt(pi) # Note: 'pi' is a predefined constant## [1] 10
## [1] 2
## [1] 1.772454To list the defined operators used by R, simply type
?Syntax into the console. This includes the arithmetic
operators
- addition
+and subtraction- - multiplication
*and division/ - exponentiation
^or**
as well as logical operators
- less than
</ greater than> - less than or equal to
<=/ greater than equal to>= - equal to
== - not equal to
!= - or
| - and
&
The concept of logical operators is to allow a program to make a
decision based on (multiple) conditions. A condition can be evaluated to
be either TRUE or FALSE (see
?Logic). For example:
2 > 5## [1] FALSEThey are often used in combination with control statements such as
the conditional if...else statement which are mentioned
below. But first back to basic functions: R also comes with a whole set
of mathematical functions including
- trigonometric functions
sin(),cos()andtan() - exponential function
exp() - statistical metrics like
min(),max(),median()
to name just a very few of them. Functions in R provide information
on how to use them. If you need help with a function, type in
help(name_of_function) or simply
?name_of_function to display the correct usage and syntax
of a function.
Objects
Everything in R we assign to a variable is an object. We can assign
objects in R with the <- operator (i.e. an arrow),
e.g. x <- 2. Commonly, other languages use
= which also works in R, however it is definitely good
practice to use the <- operator in R.
# Assign object 'x' a value of 2
x <- 2
# <- assigns the value, but does not print the result to console
# Assign another variable 'y' ...
y <- 5
# Store the product of x and y in a new object 'z'
z <- x*y
# Print z
print(z)## [1] 10Assigned variables are stored in the environment or
workspace of our session and can be listed by calling the
ls() function.
Data Types and Data Structures
The basic data types in R comprise:
- Character (or String):
sensor <- 'Landsat-8' - Integer:
n_bands <- 11L(theLtells R it is an integer, otherwise it is interpreted as numeric) - Numeric (or Float):
lambda_red <- 0.662 - Logical (or Boolean):
sun_synchronous <- TRUEorsun_synchronous <- T - Factors:
agency <- factor('NASA',levels=c('NASA','ESA','INPE'))
Understanding data types is important for a variety of reasons. For
instance, later in the course we will manipulate large rasters of
remotely sensed reflectance data which may be stored as floating point
numbers between 0 and 1. However, storing
raster matrices as floats on disk requires more memory as compared to
using an integer format. It can therefore be handy to transform data
into integer prior to saving it to disk.
For now, let us just keep in mind that there are different data types and if we now want to store not just a single value into a variable but a whole set of data, we need to undertstand the different data structures available in R:
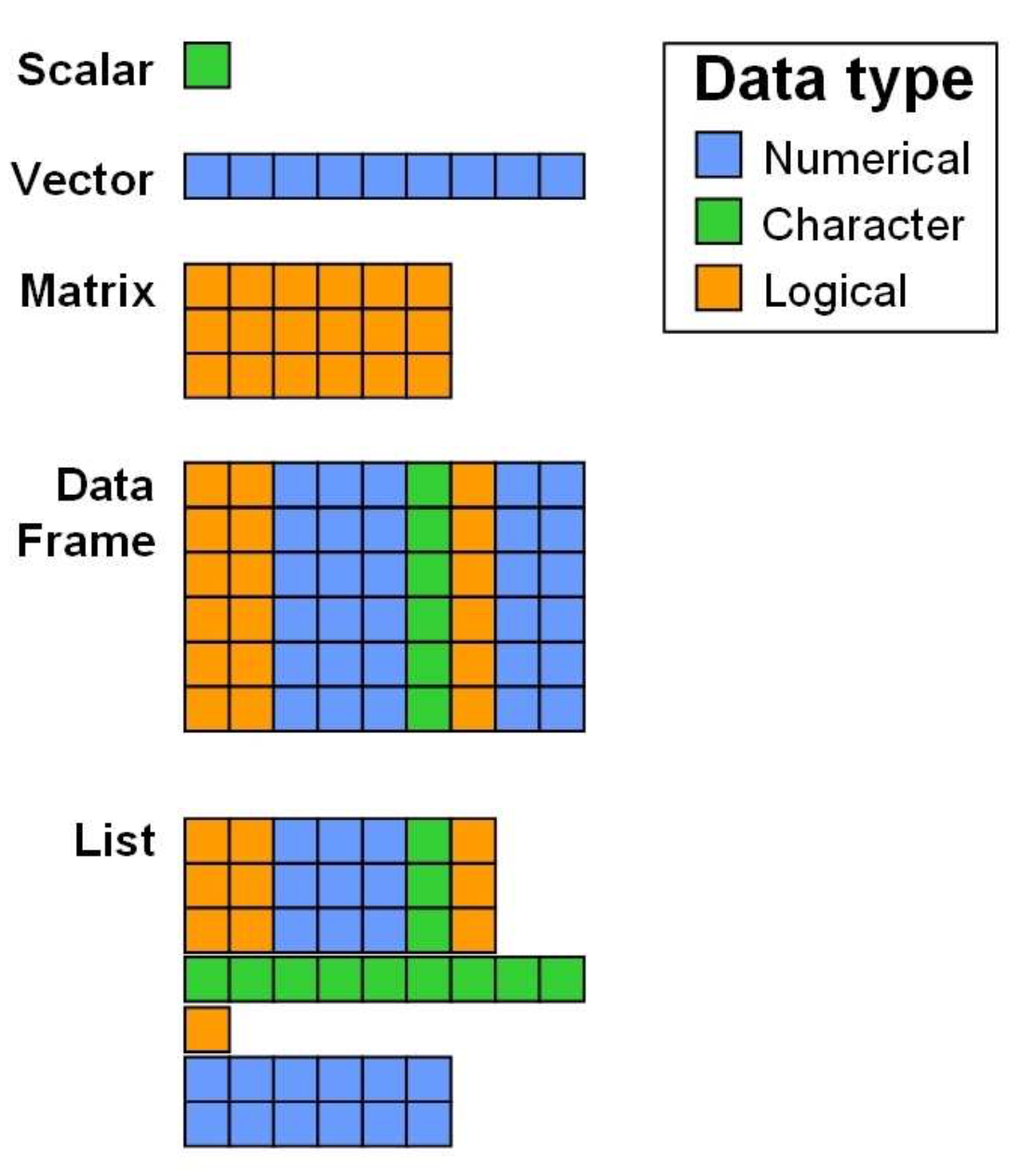
As you can see in the figure above, a variety of data structures
exist in R, which either allow for storing different data types
simultaneously (data frames, lists), or only
one datatype (scalars, vectors,
matrices). Note, that the colors representing the data
types are only chosen for vizualisation purposes: It does not mean that
matrices or vectors can only store values of type logical or numeric,
but only data of the same type.
Vectors
A vector in R can be created using a variety of functions. The most
basic function is the c()-function, where ‘c’ stands for
concatenate.
# Create a vector of numbers
a <- c(2, 4, 8, 16, 32)
# Print vector a in console
print(a)
# cat-function: Similar to print, outputs the objects, concatenating the representations
cat('vector a:', a) ## [1] 2 4 8 16 32
## vector a: 2 4 8 16 32Let`s see what happens when we mix numeric values with strings and combine them into a vector.
b <- c(2, 4, 8, 16, 32, 'Landsat', 'Sentinel-2')
print(typeof(b))## [1] "character"Our vector was automatically cast to character. This is
because R will try and convert a smaller data type into a larger one to
avoid data loss (here from numeric to
character). The strings (Landsat and
Sentinel-2) in vector b cannot be cast to
numeric, however we can very well transform a numeric 2
into a string '2'. Be aware of this behaviour and use other
data structures if needed.
Back to vector creation. Instead of having to type in sequences or
replicates manually by hand, R provides the functions seq()
and rep().
# Create a numeric sequence and store in object s
s <- seq(from=1, to=365, by=1) # or simply s <- seq(1, 365, 1)
cat('Length of s:', length(s))
# Alternatively use (always with by=1):
s <- c(1:365)
# We may want to repeat a value n times...
r <- rep(1, 5)
print(r)
# ...or repeat a sequence
r <- rep(seq(2, 8, 2), 3)
print(r)
# ...or repeat each entry in a sequence
r <- rep(seq(2, 8, 2), each=3)
print(r)## Length of s: 365[1] 1 1 1 1 1
## [1] 2 4 6 8 2 4 6 8 2 4 6 8
## [1] 2 2 2 4 4 4 6 6 6 8 8 8Matrices and Arrays
Creating matrices and arrays is just as easy as creating vectors.
Matrices are just 2-dimensional arrays with a specified number of
rows and columns. All columns in a matrix must
have the same data type (e.g., numeric, character) and must be of equal
length.
# Creates empty matrix of size 5x2
m1 <- matrix(data=NA, nrow=5, ncol=2)
print(m1)
# Creates matrix of size 5x2 containing a numeric vector
m2<- matrix(data=c(1:10), nrow=5, ncol=2)
print(m2)
# Can you describe what byrow=T does?
m3 <- matrix(data=c(1:10), nrow=5, ncol=2, byrow = T)
print(m3)## [,1] [,2]
## [1,] NA NA
## [2,] NA NA
## [3,] NA NA
## [4,] NA NA
## [5,] NA NA
## [,1] [,2]
## [1,] 1 6
## [2,] 2 7
## [3,] 3 8
## [4,] 4 9
## [5,] 5 10
## [,1] [,2]
## [1,] 1 2
## [2,] 3 4
## [3,] 5 6
## [4,] 7 8
## [5,] 9 10Arrays are similar to matrices, but can have more than two
dimensions, as specified in the dim argument.
# Create array of two 4x3 matrices
a <- array(data=c(1:12), dim=c(4,3,2))
print(a)## , , 1
##
## [,1] [,2] [,3]
## [1,] 1 5 9
## [2,] 2 6 10
## [3,] 3 7 11
## [4,] 4 8 12
##
## , , 2
##
## [,1] [,2] [,3]
## [1,] 1 5 9
## [2,] 2 6 10
## [3,] 3 7 11
## [4,] 4 8 12In R, basic mathematical operations may be used to manipulate arrays. This can be used to add, subtract, multiply or divide a constant from every element in the array.
# Create array of one 4x3 matrices
a <- array(data=c(1:9), dim=c(4,3,1))
a2 <- a * 10We can also use these operations on multiple arrays of the same size. This array arithmetic works element-wise. For example multiplying two arrays together, the first element element of the first array is multiplied by the first element of the second array, and so on. Note: while not needed in this course, R is also capable of performing matrix algebra using these operators.
# Create two arrays each of one 4x3 matrices
a1 <- array(data=c(1:9), dim=c(4,3,1))
a2 <- array(data=c(1:9), dim=c(4,3,1))
a3 <- a1 * a2Matrices are very important structures when working with remote
sensing data, as this is how images are represented numerically. We will
extensively make use of the raster package throughout the
course where a lot of the handling is done for us, however sometimes we
need to manipulate image data with more advanced techniques and it is
therefore crucial to understand the underlying data structure.
Data Frames and Lists
Generally speaking, a data.frame is a list of vectors of
equal length, which can have varying data types.
# Create data.frame with four columns of different data types
df <- data.frame(sensor = c('Landsat-5 TM', 'Landsat-7 ETM+', 'Landsat-8 OLI-TIRS'),
n_bands = c(7, 8, 11),
active = c(F, T, T),
launched = c(1984, 1999, 2013))
print(df)## sensor n_bands active launched
## 1 Landsat-5 TM 7 FALSE 1984
## 2 Landsat-7 ETM+ 8 TRUE 1999
## 3 Landsat-8 OLI-TIRS 11 TRUE 2013The top line of the data.frame is the header, describing
the column names. Each entry is called a cell and may be indexed
(accessed) individually. We will see how this works in a moment.
Lastly, let us have a look at lists. Lists are objects that can
contain elements of different types – vectors, string scalars, matrices,
functions or yet another list. Hence, a list is a generic vector
containing other objects. The dimensions of the elements to be included
must not be identical, which gives us additional flexibility to
integrate heterogeneous datasets. A list can be created using the
list() function:
# Store the matrix, array and vector in a list
l <- list(m3, a, df)
print(l)## [[1]]
## [,1] [,2]
## [1,] 1 2
## [2,] 3 4
## [3,] 5 6
## [4,] 7 8
## [5,] 9 10
##
## [[2]]
## , , 1
##
## [,1] [,2] [,3]
## [1,] 1 5 9
## [2,] 2 6 1
## [3,] 3 7 2
## [4,] 4 8 3
##
##
## [[3]]
## sensor n_bands active launched
## 1 Landsat-5 TM 7 FALSE 1984
## 2 Landsat-7 ETM+ 8 TRUE 1999
## 3 Landsat-8 OLI-TIRS 11 TRUE 2013As the print results suggests, our three objects are now
stored at locations [[1]], [[2]] and
[[3]]. We may want to access these list elements and cells
therein individually. This brings us to the next topic - indexing.
Indexing
So how do we access values in vectors,
data.frames or the elements in the list we
just created? Each element is assigned an index (i.e. a positive
integer) which we can retrieve by addressing these indices. This is
achieved using square brackets and a given index location(s).
b <- c(2, 4, 8, 16, 32, 64, 128, 256)
# Retrieve the 4th value of b
b[4]
# Retrieve value 1, 2, and 3
b[1:3] ## [1] 16
## [1] 2 4 8Use a vector to select certain positions:
# Retrieve value 1, 3, 5, and 7 of b
b[c(1, 3, 5, 7)] ## [1] 2 8 32 128Manipulate entries:
# Set the first value in b to 0
b[1] <- 0
print(b)## [1] 0 4 8 16 32 64 128 256For lists, one generally uses [[ to select any single
element, whereas [ returns a list of the selected
elements.
# [[]] returns element 1 in its original data type
typeof(l[[1]])
# [] returns a list containing element 1
typeof(l[1])
# Create a new object containing element 1
object_l <- l[[1]]
# Create a new object containing cell [4,2] of element 1
cell_l <- l[[1]][4,2] ## [1] "integer"
## [1] "list"We may also use the $ operator to access specific
elements (in lists) or columns (in data frames), given that names were
assigned. For example:
names(l) # Currently, our list does not have names...
names(l) <- c('matrix', 'array', 'df') #... names() can be used to assign element names
print(l$df)
print(df$n_bands)## NULL
## sensor n_bands active launched
## 1 Landsat-5 TM 7 FALSE 1984
## 2 Landsat-7 ETM+ 8 TRUE 1999
## 3 Landsat-8 OLI-TIRS 11 TRUE 2013
## [1] 7 8 11If you are uncertain about the characteristics of an object, you may use additional functions to investigate its attributes. Good examples are:
str(): structure of an objectlength(): number of elements in one-dimensional objects or listsnrow(),ncol(): number of rows or columns in 2-dimensional objects (df or matrix)class(),typeof(): class or type of an objectnames(): retrieve column names of df, or element names of list
Missing Values
Often our data has missing entries. For illustration, think of a
vector of temperature measurements where due to measurement error we
miss an entry now and then. For completeness, those are not just
excluded but may often be represented by a certain value, e.g. -9999. In
R missing values can be set to the logical constant NA (Not
Available).
Let us consider an vector of hourly temperature measurements where -9999 in fact was used to represent erroneous measurements:
t_hourly <- c(-1.2, -2.3, -2.4, -2.6, -1.8, -9999, 0.2, 1.4, 2.5, 4.7, -9999, 9.9, 12.1, 13.1, 13.0, 11.8, 9.8, 8.4, 7.5, 6.2, 5.0, 4.1, 4.2, 3.8)
# Calculate daily mean temperature
mean(t_hourly)## [1] -828.775These measurements exhibit two invalid temperature values of
-9999. Accordingly, the calculated mean is not of value to
us. We need to flag these entries as invalid or more generally not
available NA:
# Use indexing with logical condition
t_hourly[t_hourly == -9999] <- NA
# mean() will yield NA as soon as one NA value appears unless...
mean(t_hourly)
# ... we specify na.rm = True to ignore missing values:
mean(t_hourly, na.rm = T)## [1] NA
## [1] 4.881818There are often various ways to achieve what we want, the best way largely depends on how we want to proceed with our data. Here are more examples to do the same:
t_hourly <- c(-1.2, -2.3, -2.4, -2.6, -1.8, -9999, 0.2, 1.4, 2.5, 4.7, -9999, 9.9, 12.1, 13.1, 13.0, 11.8, 9.8, 8.4, 7.5, 6.2, 5.0, 4.1, 4.2, 3.8)
# Use only the entries that are not -9999...
mean(t_hourly[t_hourly != -9999])
# ...or use non-NAs after assigning NA to -9999; !is.na() flags all non-NA entries
t_hourly[t_hourly == -9999] <- NA
mean(t_hourly[!is.na(t_hourly)])## [1] 4.881818
## [1] 4.881818Flow Control
Often we want code to be executed only when certain conditions are met (if/conditional-statements) or repeat a certain chain of commands in order (loops/repeating operations).
Loops
for loops can be used to iterate over items and
repeatingly execute code blocks. For example, we can loop over each item
in our hourly temperature vector and print the recorded value:
# Iterate over each entry in v and print
for (item in t_hourly){
print(item)
}## [1] -1.2
## [1] -2.3
## [1] -2.4
## [1] -2.6
## [1] -1.8
## [1] NA
## [1] 0.2
## [1] 1.4
## [1] 2.5
## [1] 4.7
## [1] NA
## [1] 9.9
## [1] 12.1
## [1] 13.1
## [1] 13
## [1] 11.8
## [1] 9.8
## [1] 8.4
## [1] 7.5
## [1] 6.2
## [1] 5
## [1] 4.1
## [1] 4.2
## [1] 3.8while repeatedly evaluates a condition and executes
commands until the final condition is fulfilled:
# Assign integer to v
v <- -3
# Adds +1 to v and then prints until v > 2
while (v <= 2) {
v <- v + 1
print(v)
}## [1] -2
## [1] -1
## [1] 0
## [1] 1
## [1] 2
## [1] 3Note that loops should be avoided in higher-level languages such as R or Python where possible because they tend to make the programs run slower than when using in-built vectorized functions. For example, calculating the sum over a vector may be calculated as follows:
s <- seq(1, 10, 0.5)
s_sum <- 0
for (i in s){
s_sum <- s_sum + i
}
print(s_sum)## [1] 104.5Much more elegant and faster (think of manipulating large datasets) is to use
s_sum <- sum(s)
print(s_sum)## [1] 104.5if-statements
For conditional statements, the most commonly used approach are
if-statements which are based on boolean logic. It is
evaluated if a condition is TRUE or FALSE and
the following code block will therefore be executed (TRUE)
or not (FALSE).
For example, let us check if there are temperatures in our hourly temperature vector which are below zero degrees celsius and record the evaluation (TRUE/FALSE) in a new vector:
# create new empty vector in which we can 'fill' if the temperatures where below 0
freezing <- numeric(0)
# now we loop over the items and check the recorded temperatures
# we need to be aware of nodata values (NAs) and check for them as well
for (t in t_hourly) {
if (is.na(t)) {
freezing <- c(freezing, NA) # append NA
} else if (t < 0){
freezing <- c(freezing, 1) # append 1 (= it is below 0)
} else {
freezing <- c(freezing, 0) # append 0 (= it is not below 0)
}
}There is also a vectorized version ifelse() which allows
us to the above in one line of code:
freezing <- ifelse(t_hourly < 0 & !is.na(t_hourly), 1, 0) # ifelse(condition, TRUE, FALSE)
print(freezing)## [1] 1 1 1 1 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0Functions
R allows for user-specified functions which comes in handy whenever a certain computation needs to be repeated or generalized. Generally, a function is a set of statements organized together to perform a specific task. Using functions greatly enhances code structure as it improves the clarity and readability of your script.
Function arguments are provided in round () brackets.
Between {} stands the function body which contains the
collection of statements. return() is used to define the
output of a function. A function may be as simple as a print statement
without any objects to be returned.
A function object is defined with the function()
call:
# Basic function
printer <- function(){
print('Hello World!')
}
printer()## [1] "Hello World!"Now we need to convert our degree celsius data to degrees fahrenheit for our non-metric american collegues. We cannot remember the conversion factors and simply want a little function in which we can plug-in our records and return the values:
# Function to convert degree C to degree F
convert_temp <- function(x, C_to_F = TRUE) {
if (C_to_F) {
x_converted <- x * (9/5) + 32
} else {
x_converted <- (x - 32) * (5/9)
}
return(x_converted)
}
# 'convert_temp' is now an R object (in this case a function) which we can use like any other function:
t_hourly_F <- convert_temp(t_hourly)
print(t_hourly_F)## [1] 29.84 27.86 27.68 27.32 28.76 NA 32.36 34.52 36.50 40.46 NA 49.82 53.78 55.58 55.40 53.24 49.64 47.12 45.50 43.16 41.00 39.38 39.56 38.84Here, x is an obligatory argument that the user has to
specify while C_to_F is optional as such that it has a
default value (TRUE) which may or may not be changed
(e.g. here be set to FALSE to convert from F to C
instead).
Data Types
The binary system represents numbers through sequences of bits (binary digits). Each bit has a position and a state. The positions are numbered from 0 to the total number of bits, usually read from right to left. The state of an individual bit can either be 0 (FALSE) or 1 (TRUE). From an electric circuits perspective, electricity is either off (0) or on (1). Consider the meaning of 0 as “FALSE” or “NO”, 1 equals “TRUE” or “YES”.
This is the very fundamental logic on which all software is currently
built upon. This is because we can represent all sort of concepts in
binary, including - of course - numbers and letters. A number in binary
is a number expressed in the base-2 numeral system (= 0
and 1, instead of 0, 1, 2, 3, 4, 5, 6, 7, 8, 9 in decimal). Just like in
the decimal system, however, if we want to represent quantities beyond
the amount of unique digits available (0-1 for binary, 0-9 for decimal),
we ‘flip over’ to an additional position, re-starting the count on the
previous position (in decimal this would be
1, 2, ..., 9, -> flip -> 10, 11, ..., 99 -> flip -> 100, 101, ..., 999 -> flip -> 1000, ...).
Hence if we run out of symbols, we add a new position.
Let us image we start counting from 0 to 10 in binary:
- 0
-> 0 - 1
-> 1- now we already run out of symbols, so for the next count we add a position - 2
-> 10 - 3
-> 11- again, we add a position for the next count - 4
-> 100 - 5
-> 101 - 6
-> 110 - 7
-> 111- again, we add a position for the next count - 8
-> 1000 - …
Since a binary system operates in base 2, each bit position is assigned a unique value defined by 2n, where n is the bit position starting at 0. So for bit position 0, we have a value of 20=1, for bit position 1 we have a value of 21=2, for bit position 2 we have a value of 22=4, and so on. When extracting values from the entire bit sequence, we can simply add together all bit positions which have a state of 1 (TRUE). The visualization of the described looks as follows:
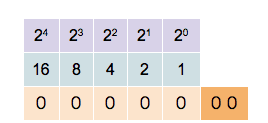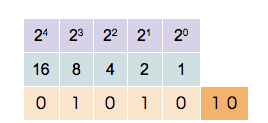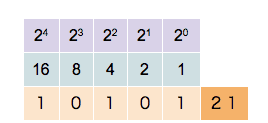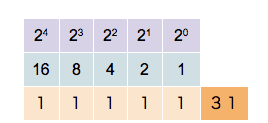
In the example above, five bit positions are given, hence the quantization is 5-bit. We can see that different combinations of states and positions enable us to represent any integer number, limited only by the number of bits in the sequence.
The more bits in a sequence we have, the higher the numbers we can generate. With 5 bits, we can represent the numbers 0–31 (i.e., 32 unique values), with 16 bits, we can represent the numbers 0 - 65,535 (i.e., 65,536 unique values).
Common data types of digital remote sensing imagery
| Data type (name) | Typical bit length | Value range | R datatype |
|---|---|---|---|
| byte / int8 | 1 byte = 8 bit | -128 to 127 (signed); 0 to 255 (unsigned) | 'INT1S'; 'INT1U' |
| int16 | 2 byte = 16 bit | -32768 to 32767 (signed); 0 to 65535 (unsigned) | 'INT2S'; 'INT2U' |
| float | 4 byte = 32 bit | -3,4e^38 to 3,4e^38, ‘single precision’; floating point number according to IEEE 754 | 'FLT4S' |
| double | 8 byte = 64 bit | -Inf to +Inf, ‘double precision’ | 'FLT8S' |
The data types byte and int16 store whole numbers while float and double store floating-point values, i.e., values that have potential decimal places. For instance, if you wanted to store the number 3.14 in integer format, it would be in the potential range of values for both types, but the decimal positions would be truncated and therefore only a value of 3 would be stored.
Now you may ask why we would bother about choosing the datatype at all, if there are obvious choices such as float / double that can capture a large range of unique values and also allow for decimal storage? Well, a greater bit length (8bit vs. 32bit) also means we need to store more information on our data storage device, possibly without needing to, because the information could be stored more efficiently with a smaller bit length. The file size or memory needed to store an image can be calculated as follows:
file size = number-of-bands × number-of-columns × numbers-of-rows × bit-length
Tutorials
We here covered only very basic aspects of R. At this stage, we did
not cover advanced data wrangling using packages such as
tidyr or dplyr, or look into advanced plotting
with ggplot2. Please follow one or several tutorials to
revisit and amend your knowledge:
General tutorials:
Graphics & visualization:
Geodata processing:
- Wageningen University: Geoscripting
- University of Colorado: EarthDataScience)
- RSpatial: Remote Sensing Image Analysis
Coding style
A few basic rules apply to coding in R. Here is a short summary of Hadley Wickham´s style guide:
Regularly save your progress.
Script names should be meaningful and end in ‘.R’.
Comment (#) your code & separate it into readable chunks.
Try to limit your code to 80 characters per line.
Variable and function names should be lowercase.
Variable names should be nouns and function names verbs.
Place spaces around operators (=, +, -, <-, etc.) and after commas.
Use <-, not =, for assignment.
An example:
######################################################
# Creating random data and a correlated response
# Author, 2020
# Load all required packages
library(ggplot2)
# Create random data
x <- runif(50, 0, 2)
# Build function to simulate response
create.response <- function(x){x + rnorm(50, 0, 0.2)}
# Apply function to random data
y <- create.response(x)
# Make a dataframe
data <- data.frame('x' = x, 'y' = y)
# Plot the simulated dataset
ggplot(data, aes(x = x, y = y)) +
geom_point()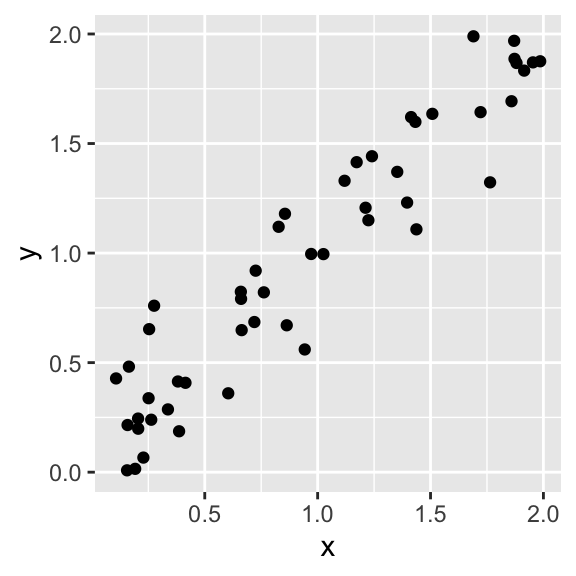
# Investigate correlation in the data
cor(data$x, data$y)## [1] 0.9517947Help!
If you get stuck while programming, there are plenty of things you can do:
- Seek the function´s help page (i.e. highlight the function and hit F1)
- Search your problem or error message
- Ask your colleagues
- Use the moodle course forum
- Check forums (e.g., StackOverflow)
Setting up a project in RStudio
We recommend to create an RStudio project for all the work you do for
this module. This helps you keep your data and scripts (.R-
or .Rmd-files) organised.
In order to set up a new project in Rstudio:
- Navigate to the tab “File” and select “New Project”
- Either select “New Directory” if you do not have a folder for this module yet, or select “Existing Directory” to point towards to such an existing folder. If (1): “New Directory” > “New Project”, select a name and a location to store the folder in, and finish by “Create Project”.
- Navigate to the path of the project folder on your machine
- Inside the folder, create the following sub-folders: data, scripts, models, results, and figures
- All scripts for each session are ideally saved into separate
.R- or.Rmd-files and saved in your “scripts” folder - You can use this blueprint .Rmd for your work
If you close and reopen RStudio, the last project should be reloaded automatically. If not the case, just navigate to the tab “File” and select “Open Project”.
Test for Session 3: terra-package
With your R console opened in RStudio, you can install the
terra package like any other package in R as follows:
# install the terra package
install.packages('terra')
# load the package to get access to the package's routines/functions in your environment
library(terra)Once the package is loaded into your current environment
(library(terra)), navigate to the Landsat-8 image you
worked with before, copy the absolute path to the .tif-file
and try to load the image into R. This should result in printing some
image properties to your console like shown below.
# create variable which contains the filepath
file_landsat8 <- "your/path/to/the/landsat8/file.tif"
# use the filepath to read in the file into a "rast()"-object
landsat_8 <- rast(file_landsat8)
print(landsat_8)## class : SpatRaster
## dimensions : 1840, 2171, 6 (nrow, ncol, nlyr)
## resolution : 30.00639, 30.00728 (x, y)
## extent : 360793.1, 425937, 5794470, 5849683 (xmin, xmax, ymin, ymax)
## coord. ref. : WGS 84 / UTM zone 33N (EPSG:32633)
## source : LC08_L1TP_193023_20190726_20200827_02_T1_int16.tif
## names : BLU, GRN, RED, NIR, SW1, SW2
## min values : 639, 371, 227, 60, 23, 16
## max values : 5603, 5984, 6728, 7454, 11152, 8946Copyright © 2023 Humboldt-Universität zu Berlin. Department of Geography.